PDB File Format

The Protein Data Bank (PDB) is a database for the three-dimensional structural data of atomic compounds, such as proteins and nucleic acids. The data, typically obtained by X-ray crystallography, NMR spectroscopy, or, increasingly, cryo-electron microscopy, and submitted by researchers from around the world, are freely accessible on the Internet via the websites of its member organizations (PDBe, PDBj, RCSB, and BMRB). The PDB is overseen by an organization called the Worldwide Protein Data Bank, wwPDB. Technically, the PDB is a key in areas of molecular dynamics simulations. Most major scientific journals, and some funding agencies, now require scientists to submit their structure data to the PDB. Many other databases use protein structures deposited in the PDB. For example, SCOP and CATH classify protein structures, while PDBsum provides a graphic overview of PDB entries using information from other sources, such as Gene ontology. The simplified diagram of atom records in the PDB format depicted in below figure.
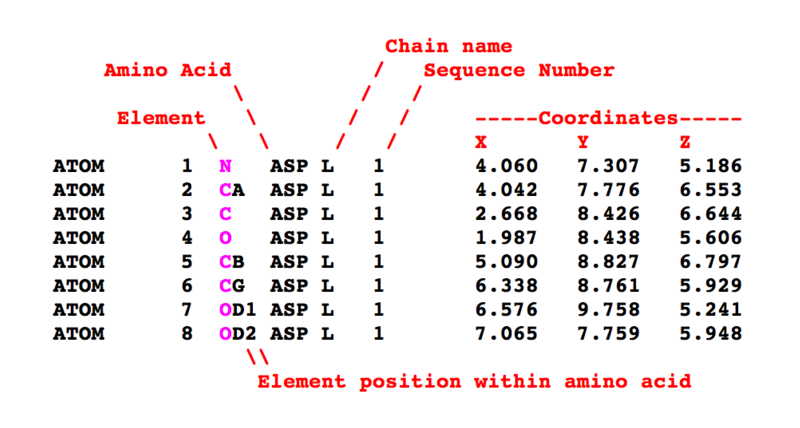
The described PDB files may be viewed using one of several free and open source softwares, including Jmol, Pymol, VMD, and Rasmol. Other non-free, shareware programs include ICM-Browser, MDL Chime, UCSF Chimera, Swiss-PDB Viewer, StarBiochem (a Java-based interactive molecular viewer with integrated search of protein databank), Sirius, and VisProt3DS (a tool for Protein Visualization in 3D stereoscopic view in anaglyth and other modes), and Discovery Studio. Furthermore, the RCSB PDB website contains an extensive list of both free and commercial molecule visualization programs and web browser plugins.
Reference
Berman, H., Henrick, K., & Nakamura, H. (2003). Announcing the worldwide Protein Data Bank. Nature Structural & Molecular Biology, 10(12), 980–980.
doi:10.1038/nsb1203-980.
