Uniaxial Tension Simulation
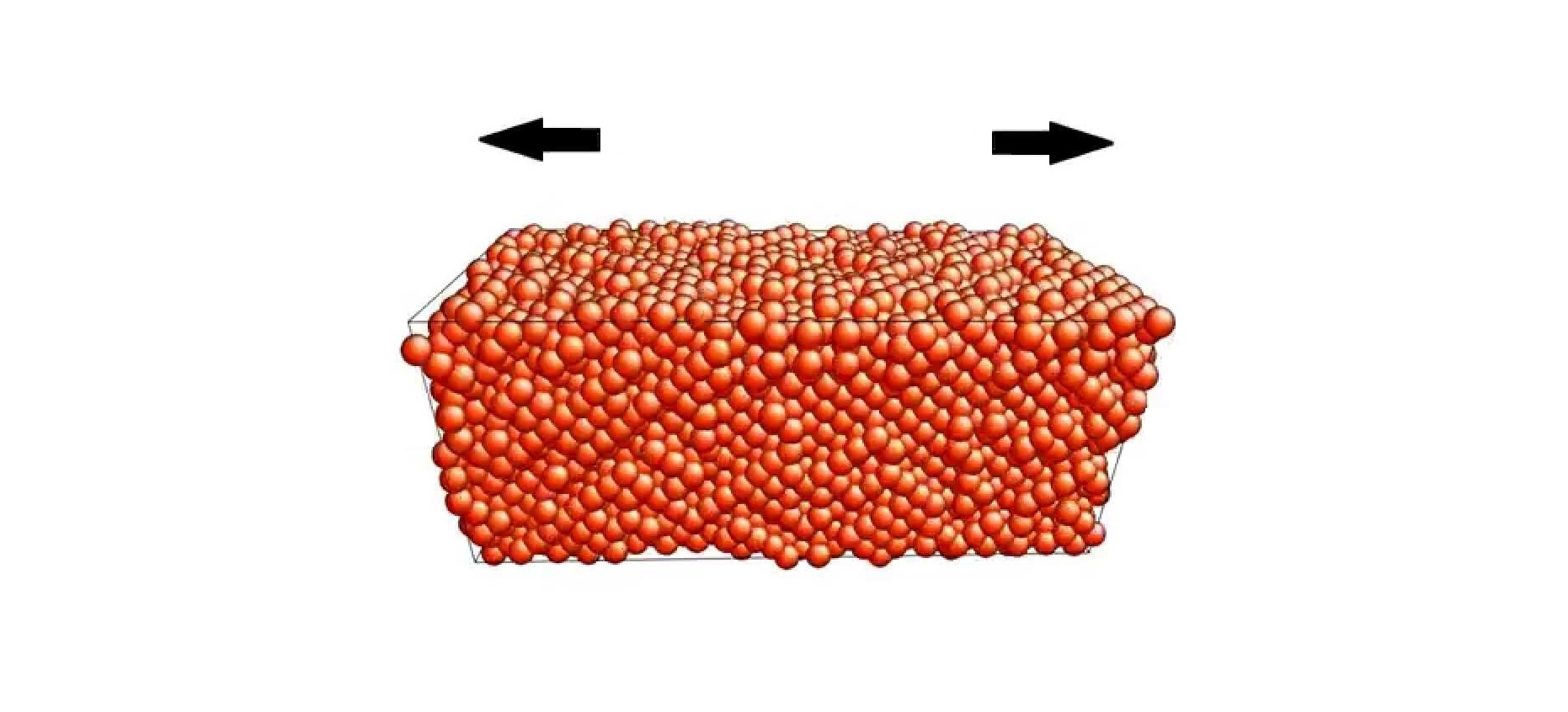
This LAMMPS script shows how to run an atomistic simulation of uniaxial tensile loading of an aluminum single crystal oriented in the <100> direction.
# Input file for uniaxial tensile loading of single crystal aluminum
# Mark Tschopp, November 2010
# INITIALIZATION
units metal
dimension 3
boundary p p p
atom_style atomic
variable latparam equal 4.05
# ATOM DEFINITION
lattice fcc ${latparam}
region whole block 0 10 0 10 0 10
create_box 1 whole
lattice fcc ${latparam} orient x 1 0 0 orient y 0 1 0 orient z 0 0 1
create_atoms 1 region whole
# FORCE FIELDS
pair_style eam/alloy
pair_coeff * * Al99.eam.alloy Al
# SETTINGS
compute csym all centro/atom fcc
compute peratom all pe/atom
# EQUILIBRATION
reset_timestep 0
timestep 0.001
velocity all create 300 12345 mom yes rot no
fix 1 all npt temp 300 300 1 iso 0 0 1 drag 1
# Set thermo output
thermo 1000
thermo_style custom step lx ly lz press pxx pyy pzz pe temp
# Run for at least 10 picosecond (assuming 1 fs timestep)
run 20000
unfix 1
# Store final cell length for strain calculations
variable tmp equal "lx"
variable L0 equal ${tmp}
print "Initial Length, L0: ${L0}"
# DEFORMATION
reset_timestep 0
fix 1 all npt temp 300 300 1 y 0 0 1 z 0 0 1 drag 1
variable srate equal 1.0e10
variable srate1 equal "v_srate / 1.0e12"
fix 2 all deform 1 x erate ${srate1} units box remap x
# Output strain and stress info to file for units metal
variable strain equal "(lx - v_L0)/v_L0"
variable p1 equal "v_strain"
variable p2 equal "-pxx/10000"
variable p3 equal "-pyy/10000"
variable p4 equal "-pzz/10000"
fix def1 all print 100 "${p1} ${p2} ${p3} ${p4}" file Al_SC_100.def1.txt screen no
# Use XYZ for VMD Software
dump 1 all cfg 250 dump.tensile_*.cfg mass type xs ys zs c_csym c_peratom fx fy fz
dump_modify 1 element Al
# Display thermo
thermo 1000
thermo_style custom step v_strain temp v_p2 v_p3 v_p4 ke pe press
run 20000
# SIMULATION DONE
print "All done"

This molecular dynamics simulation first generates a simulation cell with fcc atoms with <100> orientations in the x, y, and z-directions. For this example, the simulation cell size is 10 lattice units in each direction, i.e., 4000 total atoms. Larger simulation cell sizes should be used to converge the dislocation nucleation stress values and to not influence the dislocation nucleation mechanism. The potential used here is the Mishin et al. (1999) aluminum potential. The equilibration step allows the lattice to expand to a temperature of 300 K with a pressure of 0 bar at each simulation cell boundary. Then, the simulation cell is deformed in the x-direction at a strain rate of 10^10 1/s, while the lateral boundaries are controlled using the NPT equations of motion to zero pressure. The stress and strain values are output to a separate file, which can be imported in a graphing application for plotting. The XYZ dump files include the x, y, and z coordinates for each atom. This can be directly visualized using VMD software.
Reference
https://lammps.sandia.gov
