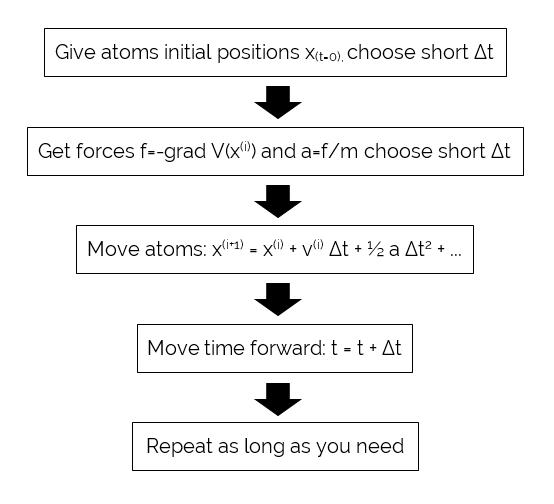
The Molecular Dynamics Process
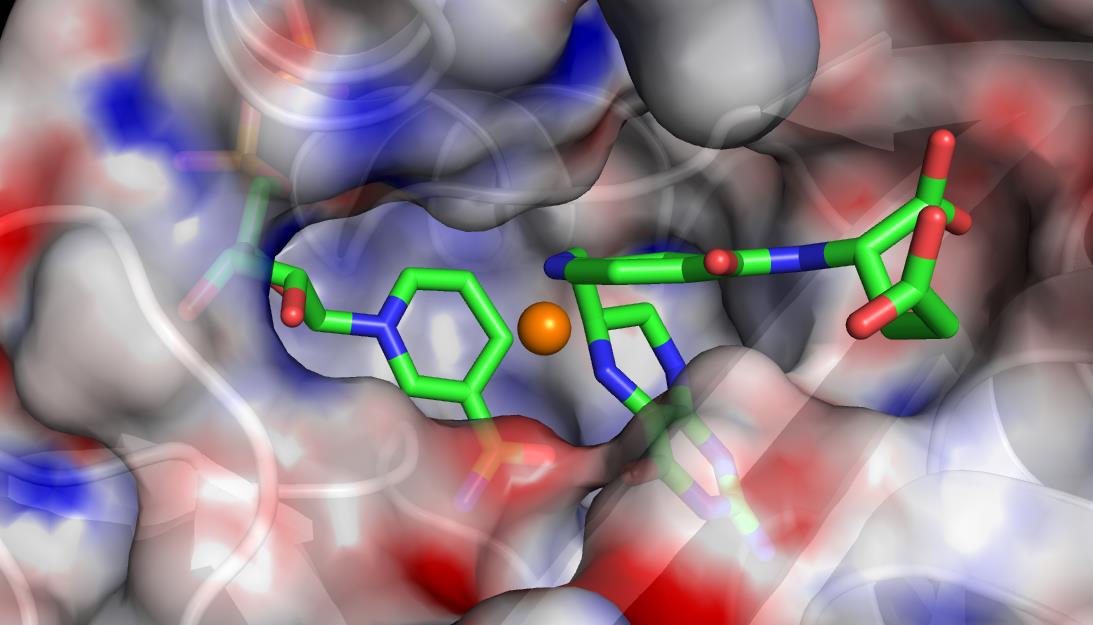
Classical molecular dynamics (MD) is a branch of computational science that focuses on simulating the motion of particles (atoms and molecules) according to Newton’s Laws of Motion. particles are approximated as “balls on springs,” which allows for the application of the following laws:
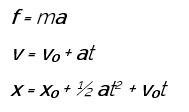
where x is position, v is velocity, a is acceleration, and t is simulation time. Classical MD approximations treat bond and angle forces as harmonic, and dihedrals as periodic. After interaction (force) definition, MD simulations are conducted in a series of time steps. In each time step, the force on each particle is calculated according to the derivative of the energy equation. Next, particle is moved a certain distance corresponding to the magnitude and direction of the force over the size of the time step. The process is repeated over many time steps, iteratively calculating forces and solving the equations of motion based on the accelerations from the forces, and produces results. A simplified view of the computational steps a MD simulation will take, depicted in below figure. Production MD codes use more complicated versions of this algorithm that incorporate temperature and pressure control.